An algorithm for partial SVD (or PCA) of a Filebacked Big Matrix through the eigen decomposition of the covariance between variables (primal) or observations (dual). Use this algorithm only if there is one dimension that is much smaller than the other. Otherwise use big_randomSVD.
big_SVD(
X,
fun.scaling = big_scale(center = FALSE, scale = FALSE),
ind.row = rows_along(X),
ind.col = cols_along(X),
k = 10,
block.size = block_size(nrow(X))
)Arguments
- X
An object of class FBM.
- fun.scaling
A function with parameters
X,ind.rowandind.col, and that returns a data.frame with$centerand$scalefor the columns corresponding toind.col, to scale each of their elements such as followed: $$\frac{X_{i,j} - center_j}{scale_j}.$$ Default doesn't use any scaling. You can also provide your owncenterandscaleby usingas_scaling_fun().- ind.row
An optional vector of the row indices that are used. If not specified, all rows are used. Don't use negative indices.
- ind.col
An optional vector of the column indices that are used. If not specified, all columns are used. Don't use negative indices.
- k
Number of singular vectors/values to compute. Default is
10. This algorithm should be used to compute only a few singular vectors/values. If more is needed, have a look at https://stackoverflow.com/a/46380540/6103040.- block.size
Maximum number of columns read at once. Default uses block_size.
Value
A named list (an S3 class "big_SVD") of
d, the singular values,u, the left singular vectors,v, the right singular vectors,center, the centering vector,scale, the scaling vector.
Note that to obtain the Principal Components, you must use predict on the result. See examples.
Details
To get \(X = U \cdot D \cdot V^T\),
if the number of observations is small, this function computes \(K_(2) = X \cdot X^T \approx U \cdot D^2 \cdot U^T\) and then \(V = X^T \cdot U \cdot D^{-1}\),
if the number of variable is small, this function computes \(K_(1) = X^T \cdot X \approx V \cdot D^2 \cdot V^T\) and then \(U = X \cdot V \cdot D^{-1}\),
if both dimensions are large, use big_randomSVD instead.
Matrix parallelization
Large matrix computations are made block-wise and won't be parallelized
in order to not have to reduce the size of these blocks. Instead, you can use
the MKL
or OpenBLAS in order to accelerate these block matrix computations.
You can control the number of cores used by these optimized matrix libraries
with bigparallelr::set_blas_ncores().
See also
Examples
set.seed(1)
X <- big_attachExtdata()
n <- nrow(X)
# Using only half of the data
ind <- sort(sample(n, n/2))
test <- big_SVD(X, fun.scaling = big_scale(), ind.row = ind)
str(test)
#> List of 5
#> $ d : num [1:10] 178.5 114.5 91 87.1 86.3 ...
#> $ u : num [1:258, 1:10] -0.1092 -0.0928 -0.0806 -0.0796 -0.1028 ...
#> $ v : num [1:4542, 1:10] 0.00607 0.00739 0.02921 -0.01283 0.01473 ...
#> $ center: num [1:4542] 1.34 1.63 1.51 1.64 1.09 ...
#> $ scale : num [1:4542] 0.665 0.551 0.631 0.55 0.708 ...
#> - attr(*, "class")= chr "big_SVD"
plot(test$u)
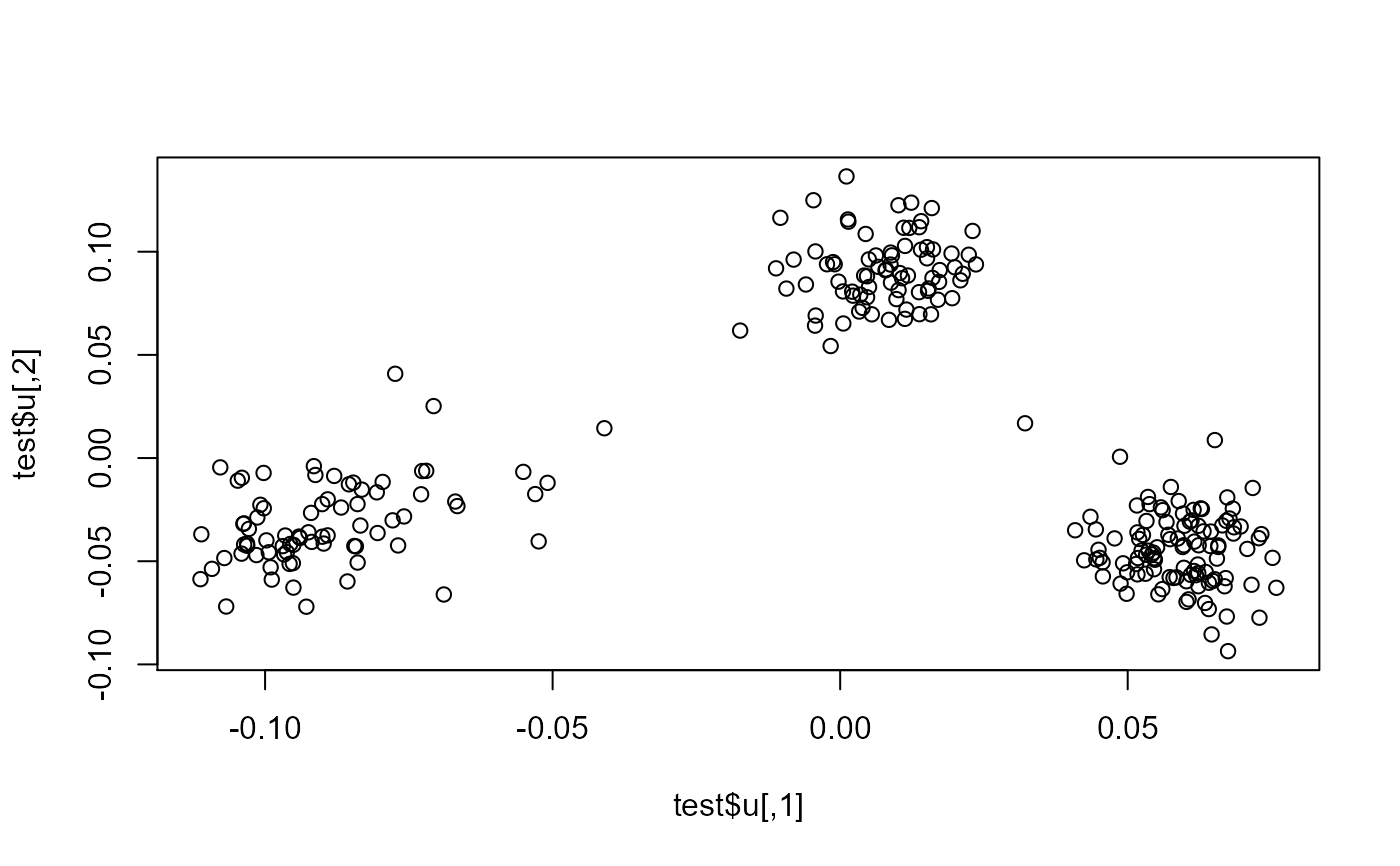 pca <- prcomp(X[ind, ], center = TRUE, scale. = TRUE)
# same scaling
all.equal(test$center, pca$center)
#> [1] TRUE
all.equal(test$scale, pca$scale)
#> [1] TRUE
# scores and loadings are the same or opposite
# except for last eigenvalue which is equal to 0
# due to centering of columns
scores <- test$u %*% diag(test$d)
class(test)
#> [1] "big_SVD"
scores2 <- predict(test) # use this function to predict scores
all.equal(scores, scores2)
#> [1] TRUE
dim(scores)
#> [1] 258 10
dim(pca$x)
#> [1] 258 258
tail(pca$sdev)
#> [1] 3.023287e+00 3.008386e+00 2.990514e+00 2.984375e+00 2.965688e+00
#> [6] 1.130391e-14
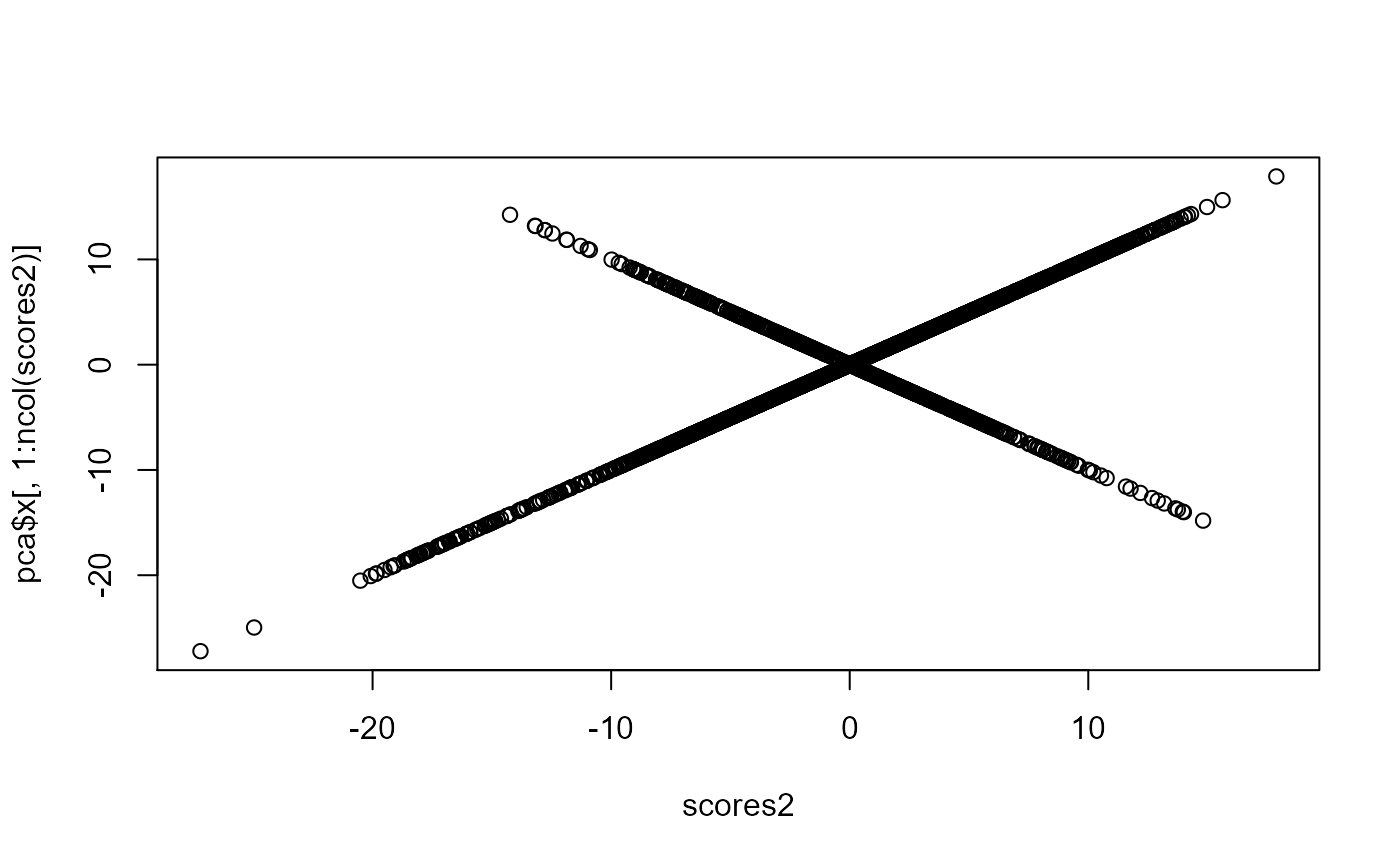
plot(scores2, pca$x[, 1:ncol(scores2)])
pca <- prcomp(X[ind, ], center = TRUE, scale. = TRUE)
# same scaling
all.equal(test$center, pca$center)
#> [1] TRUE
all.equal(test$scale, pca$scale)
#> [1] TRUE
# scores and loadings are the same or opposite
# except for last eigenvalue which is equal to 0
# due to centering of columns
scores <- test$u %*% diag(test$d)
class(test)
#> [1] "big_SVD"
scores2 <- predict(test) # use this function to predict scores
all.equal(scores, scores2)
#> [1] TRUE
dim(scores)
#> [1] 258 10
dim(pca$x)
#> [1] 258 258
tail(pca$sdev)
#> [1] 3.023287e+00 3.008386e+00 2.990514e+00 2.984375e+00 2.965688e+00
#> [6] 1.130391e-14
plot(scores2, pca$x[, 1:ncol(scores2)])
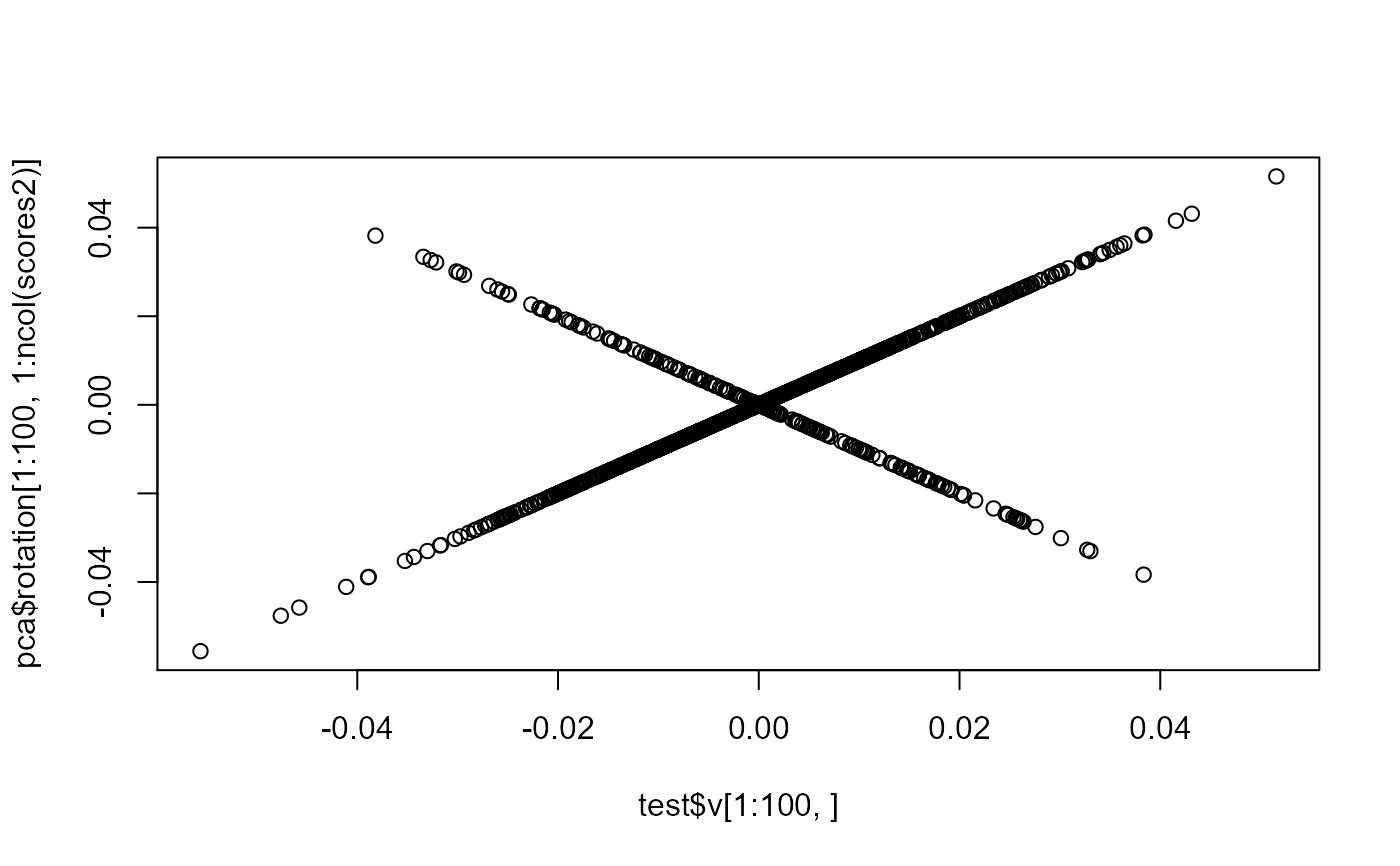
 plot(test$v[1:100, ], pca$rotation[1:100, 1:ncol(scores2)])
plot(test$v[1:100, ], pca$rotation[1:100, 1:ncol(scores2)])
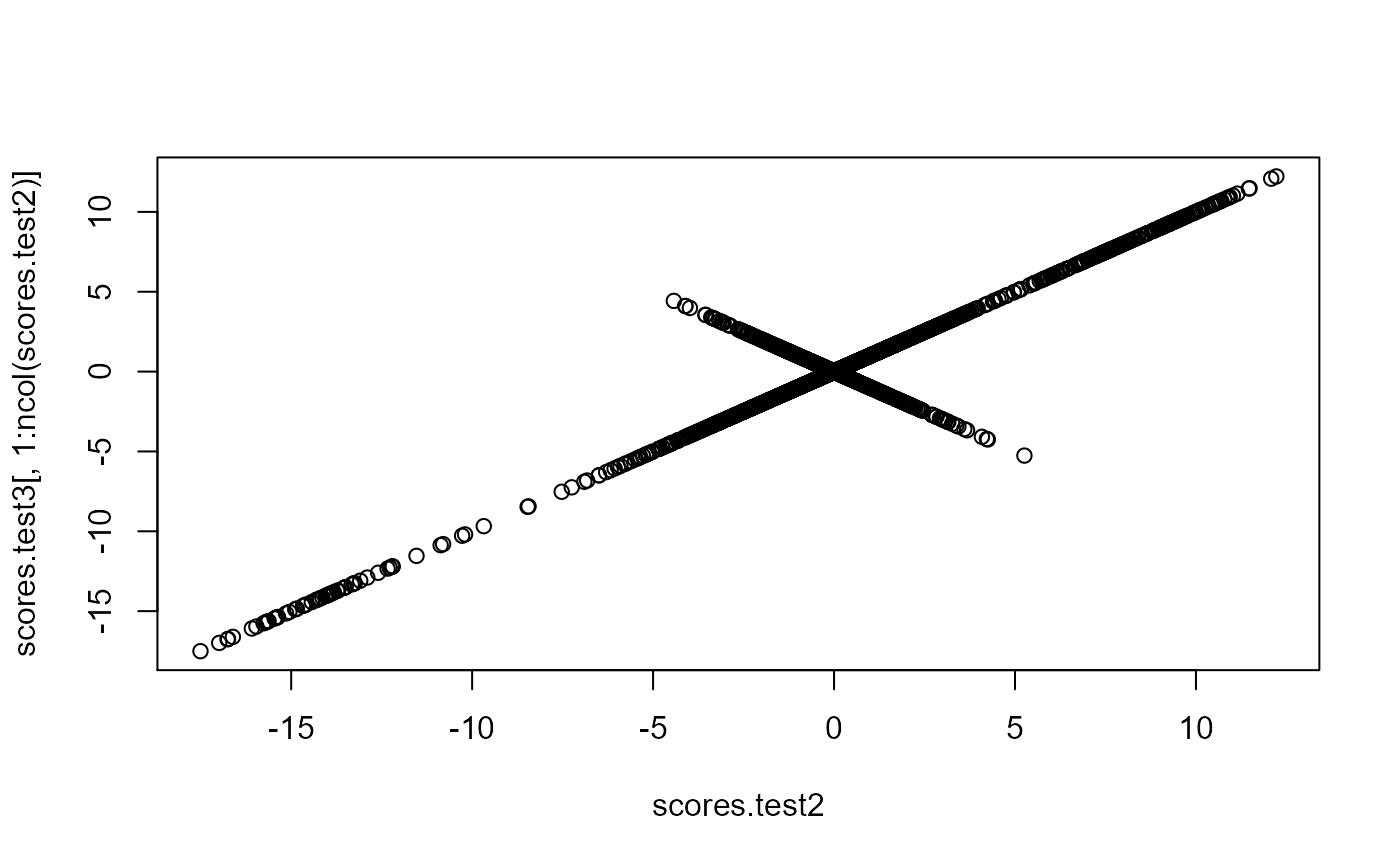
 # projecting on new data
X2 <- sweep(sweep(X[-ind, ], 2, test$center, '-'), 2, test$scale, '/')
scores.test <- X2 %*% test$v
ind2 <- setdiff(rows_along(X), ind)
scores.test2 <- predict(test, X, ind.row = ind2) # use this
all.equal(scores.test, scores.test2)
#> [1] TRUE
scores.test3 <- predict(pca, X[-ind, ])
plot(scores.test2, scores.test3[, 1:ncol(scores.test2)])
# projecting on new data
X2 <- sweep(sweep(X[-ind, ], 2, test$center, '-'), 2, test$scale, '/')
scores.test <- X2 %*% test$v
ind2 <- setdiff(rows_along(X), ind)
scores.test2 <- predict(test, X, ind.row = ind2) # use this
all.equal(scores.test, scores.test2)
#> [1] TRUE
scores.test3 <- predict(pca, X[-ind, ])
plot(scores.test2, scores.test3[, 1:ncol(scores.test2)])