Polygenic Risk Scores with possible clumping and thresholding.
Arguments
- G
A FBM.code256 (typically
<bigSNP>$genotypes).
You shouldn't have missing values. Also, remember to do quality control, e.g. some algorithms in this package won't work if you use SNPs with 0 MAF.- betas.keep
Numeric vector of weights associated with each SNP corresponding to
ind.keep. You may want to see bigstatsr::big_univLinReg or bigstatsr::big_univLogReg.- ind.test
The individuals on whom to project the scores. Default uses all.
- ind.keep
Column (SNP) indices to use (if using clumping, the output of snp_clumping). Default doesn't clump.
- same.keep
A logical vector associated with
betas.keepwhether the reference allele is the same for G. Default is allTRUE(for example when you train the betas on the same dataset). Otherwise, use same_ref.- lpS.keep
Numeric vector of
-log10(p-value)associated withbetas.keep. Default doesn't use thresholding.- thr.list
Threshold vector on
lpS.keepat which SNPs are excluded if they are not significant enough. Default doesn't use thresholding.
Value
A matrix of scores, where rows correspond to ind.test and
columns correspond to thr.list.
Examples
test <- snp_attachExtdata()
G <- big_copy(test$genotypes, ind.col = 1:1000)
CHR <- test$map$chromosome[1:1000]
POS <- test$map$physical.position[1:1000]
y01 <- test$fam$affection - 1
# PCA -> covariables
obj.svd <- snp_autoSVD(G, infos.chr = CHR, infos.pos = POS)
#> Discarding 0 variant with MAC < 10 or MAF < 0.02.
#>
#> Phase of clumping (on MAF) at r^2 > 0.2.. keep 952 variants.
#>
#> Iteration 1:
#> Computing SVD..
#> 26 outlier variants detected..
#>
#> Iteration 2:
#> Computing SVD..
#> 0 outlier variant detected..
#>
#> Converged!
# train and test set
ind.train <- sort(sample(nrow(G), 400))
ind.test <- setdiff(rows_along(G), ind.train) # 117
# GWAS
gwas.train <- big_univLogReg(G, y01.train = y01[ind.train],
ind.train = ind.train,
covar.train = obj.svd$u[ind.train, ])
# clumping
ind.keep <- snp_clumping(G, infos.chr = CHR,
ind.row = ind.train,
S = abs(gwas.train$score))
# -log10(p-values) and thresolding
summary(lpS.keep <- -predict(gwas.train)[ind.keep])
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> 0.000569 0.146282 0.344033 0.474463 0.640888 6.191733
thrs <- seq(0, 4, by = 0.5)
nb.pred <- sapply(thrs, function(thr) sum(lpS.keep > thr))
# PRS
prs <- snp_PRS(G, betas.keep = gwas.train$estim[ind.keep],
ind.test = ind.test,
ind.keep = ind.keep,
lpS.keep = lpS.keep,
thr.list = thrs)
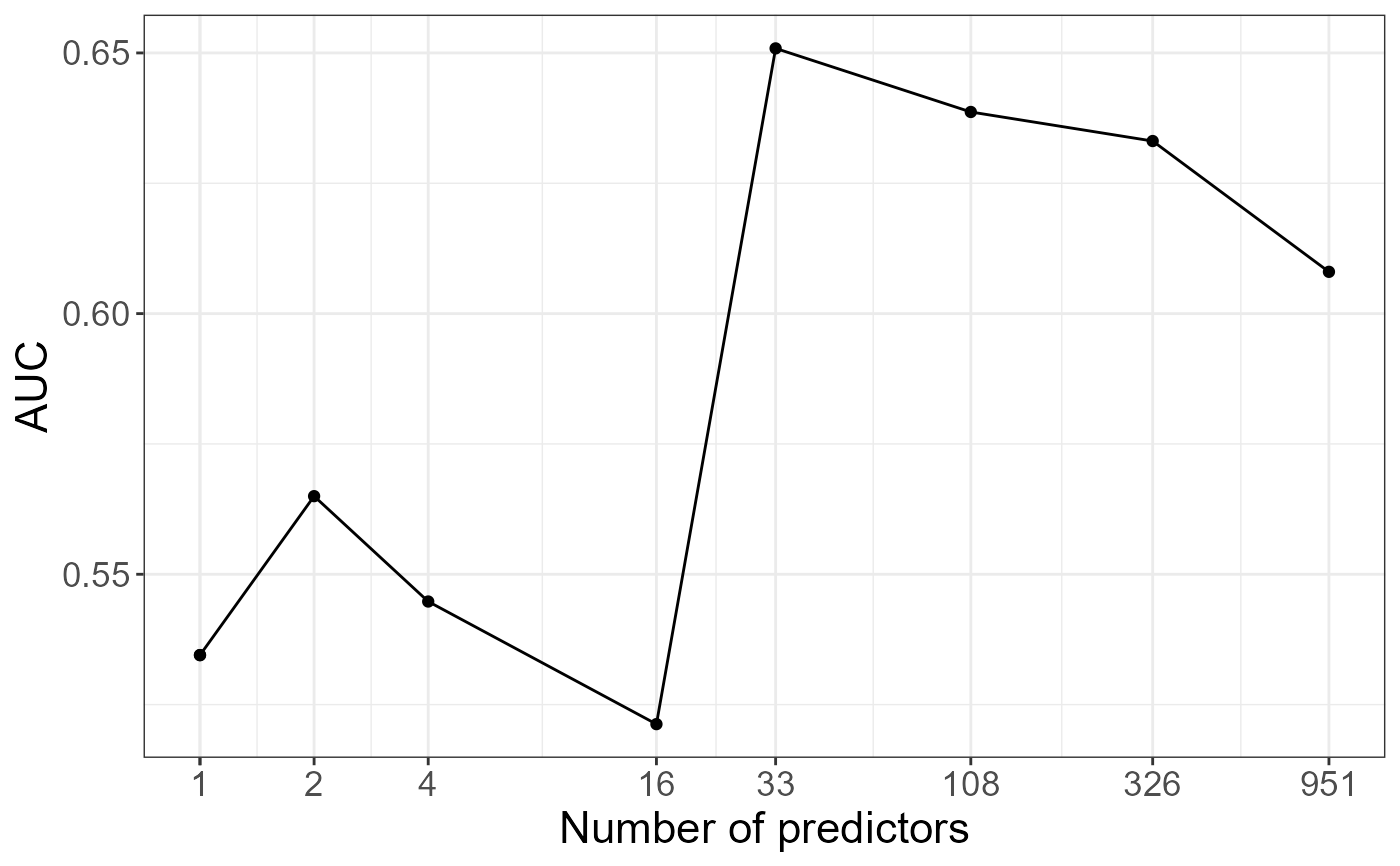
# AUC as a function of the number of predictors
aucs <- apply(prs, 2, AUC, target = y01[ind.test])
library(ggplot2)
#> Warning: package 'ggplot2' was built under R version 4.2.3
qplot(nb.pred, aucs) +
geom_line() +
scale_x_log10(breaks = nb.pred) +
labs(x = "Number of predictors", y = "AUC") +
theme_bigstatsr()
#> Warning: `qplot()` was deprecated in ggplot2 3.4.0.